Part 1 of this blog series explains how query-entity relevance spreading across a knowledge graph eliminates the need of typical iterative retrieve → reason → retrieve cycles.
Introduction
While building a GraphRAG Document Repository last year, I kept thinking about how the human brain actually retrieves information. You hear a name — a place, a person, a scientific term — and your brain doesn’t search for an exact match. Instead, that single cue triggers a cascade of associated memories: related names, connected concepts, adjacent experiences. One activation leads to the next, spreading outward through a web of associations.
That process of associative recollection struck me as conceptually similar to knowledge graph traversal — moving from one entity node to another along edges that describe how they are related. It turns out this isn’t a novel observation: knowledge graphs are a well-established tool in computational neuroscience for modeling how the hippocampus, the brain’s memory hub, encodes and retrieves associations.
That analogy became the design principle behind ENGRAM — a GraphRAG system implementing hippocampal-inspired associative retrieval using knowledge graphs and Personalized PageRank (PPR). When a user asks a question, ENGRAM extracts entities from the query and seeds them into a domain knowledge graph. PPR then propagates their relevance outward along graph edges, activating connected entities and the literature passages that cite them, – all in a single traversal, without the iterative retrieve → reason → retrieve cycles that conventional RAG systems require.
The name comes from neuroscience: an engram is the hypothetical physical substrate of memory which is a trace encoded in neural connections. ENGRAM, the system, encodes domain knowledge in graph connections and retrieves it through the same spreading-activation pattern.
In this three-part series, I’ll walk through ENGRAM’s architecture, from the core retrieval method (this post) to multi-agent response synthesis and long document support (Part 2) to the hippocampal-inspired memory system for agents (Part 3). The full codebase is available at github.com/pvelua/engram-biomed-research.
The Multi-Hop Retrieval Problem
I start with an example showing why biomedical research has been chosen as a representative use case for ENGRAM development. Biomedical researchers spend an estimated 40–60% of their time on literature review, yet frequently miss critical connections between findings scattered across separate papers. Consider a drug researcher investigating Alzheimer’s treatments who needs to discover that:
- Drug X (mentioned in Paper A) inhibits Enzyme Y
- Enzyme Y (mentioned in Paper B) regulates Protein Z
- Protein Z (mentioned in Paper C) is elevated in Alzheimer’s patients
No single paper contains all three facts. The researcher must manually chain these discoveries, or hope that a search system can do it for them.
Standard RAG systems handle this through an iterative pattern: retrieve relevant chunks, feed them to an LLM, let the LLM reason about what to search for next, retrieve again, and repeat. This retrieve → reason → retrieve cycle works, but at a cost. Each iteration requires an LLM inference call, adding latency and expense. The reasoning chain can drift off-topic, especially when intermediate results are ambiguous. And the number of iterations needed is unpredictable as the system doesn’t know in advance how many hops separate the query from the answer.
ENGRAM takes a fundamentally different approach. Instead of asking an LLM to reason about what to retrieve next, ENGRAM lets the structure of a knowledge graph do the multi-hop traversal. Query entities are seeded into the graph, and Personalized PageRank propagates their relevance outward along existing edges such as from drugs to their targets, from targets to pathways, from pathways to diseases, and from all of these to the literature passages that cite them. The entire multi-hop chain is traversed in one graph operation, with no iterative LLM calls.
Hippocampal-Inspired Associative Retrieval
The Hippocampal Analogy
The hippocampus, a structure deep in the temporal lobe, is central to how the brain forms and retrieves memories. One of its key capabilities is pattern completion — given a partial cue, the hippocampus can activate a full memory trace by spreading activation across neural associations. You smell a particular spice, and suddenly you recall a specific kitchen, a conversation, a person’s face. The partial cue (the smell) activated a network of associated memories through spreading excitation.
ENGRAM mirrors this mechanism in its retrieval design. A user’s query provides partial cues — a few entity mentions like “neuroinflammation” or “TREM2.” These cues are resolved to nodes in a knowledge graph, and Personalized PageRank spreads their relevance outward across graph edges, activating connected compounds, genes, pathways, and the literature passages that cite them. The partial cue completes into a full evidence context.
I want to be clear: this is a design analogy, not a neuroscience claim. ENGRAM doesn’t simulate hippocampal circuits. But the analogy is useful because it guides specific architectural decisions such as dual representation (vector similarity + graph structure), single-step spreading activation instead of iterative search, and the ability to complete patterns from partial cues.
The Knowledge Graph as Activation Substrate
ENGRAM’s retrieval depends on a two-layer graph structure that connects domain knowledge to the literature that references it.
The domain layer is a curated knowledge graph of biomedical entities and their relationships. For the Biomedical Research Assistant, ENGRAM uses Hetionet v1.0 — an open, integrative network of approximately 47,000 biomedical entities (genes, diseases, compounds, pathways, anatomical structures, biological processes) connected by roughly 2.25 million curated relationships (BINDS, TREATS, REGULATES, PARTICIPATES, EXPRESSES, and others). This is the “seed graph” which encodes the domain knowledge that the system expects to encounter in user queries.
The literature layer connects research text to the domain graph. PubMed abstracts (approximately 50,000, focused on Alzheimer’s and neurodegeneration) are stored as Passage nodes. User-uploaded research PDFs are processed into Article → Section → Passage hierarchies (will be covered in detail in Part 2 of this blog). The bridge between layers is formed by NounPhrase nodes: entity mentions extracted from passage text are linked to their corresponding Hetionet entities via MAPS_TO edges, with confidence scores from a two-stage resolution process (VoyageAI embedding similarity followed by ColBERT token-level re-ranking).
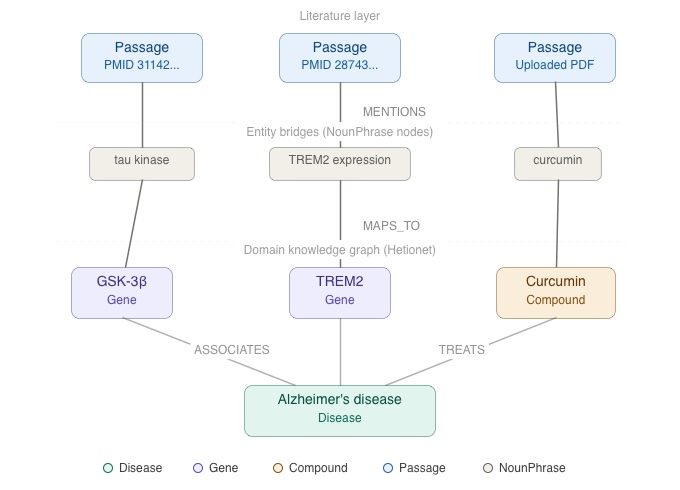
This dual-layer design means that domain relationships and literature evidence live in the same graph. When PPR spreads relevance from a disease entity, it naturally flows through gene and compound nodes *and* through the NounPhrase bridges into the passages that discuss them.
Personalized PageRank as Spreading Activation
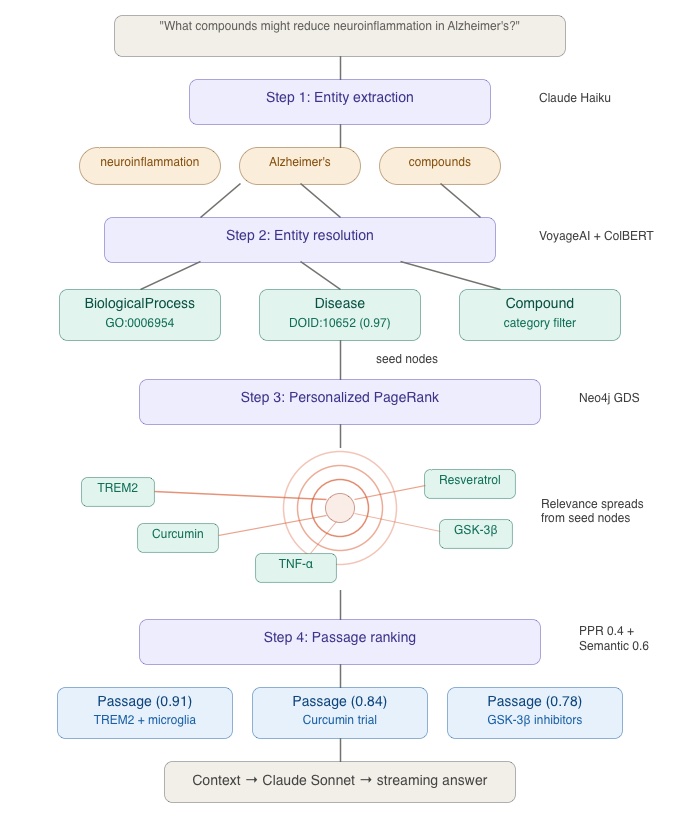
The retrieval pipeline has four steps. Let me walk you through them using a concrete example.
User Query: “What compounds might reduce neuroinflammation in Alzheimer’s?”
Step 1 — Entity extraction. An LLM (Claude Haiku) extracts named entities from the query text: `neuroinflammation`, `Alzheimer’s`, and `compounds` (as a category mention). This step identifies the “cues” that will seed the graph traversal.
Step 2 — Entity resolution. Each extracted mention is resolved to a node in the knowledge graph. “Alzheimer’s” maps to `Disease::DOID:10652` (Alzheimer’s disease in Hetionet). “Neuroinflammation” resolves to a BiologicalProcess node. Resolution uses VoyageAI embeddings for initial candidate retrieval from ChromaDB, followed by ColBERT reranking for precision. Confidence scores are attached to each resolution — a high-confidence match like “Alzheimer’s disease” (0.97) anchors the traversal, while lower-confidence matches contribute less seed weight.
Step 3 — PPR traversal. The resolved entity IDs become seed nodes for Personalized PageRank, executed via Neo4j’s Graph Data Science library. PPR works by simulating random walks that restart from the seed nodes with a fixed probability. The result is a relevance score for every reachable node in the graph — nodes closer to the seeds (in graph-hop distance) and with more paths connecting them to the seeds receive higher scores.
From our example seeds, relevance spreads outward:
Alzheimer's Disease → microglia activation → TREM2 →
→ anti-inflammatory compounds → Curcumin, Resveratrol
→ clinical trial passages citing these compounds
Neuroinflammation → cytokine signaling → TNF-α, IL-1β →
→ NounPhrases mentioning these in passage text →
→ Passage nodes in PubMed abstracts
The critical point: this entire traversal happens in one PPR computation. There is no LLM call between the disease node and the compound node, no reasoning step to decide “I should search for TREM2 next.” The graph structure *already encodes* that TREM2 is connected to neuroinflammation and Alzheimer’s. The PPR simply reveals those connections by propagating relevance scores.
Step 4 — Passage ranking. PPR assigns scores to Passage nodes based on their graph proximity to the query entities. These graph-structural scores are blended with semantic similarity scores (VoyageAI cosine distance between the query text and each passage embedding) to produce a final ranking. The blend weights — 40% PPR, 60% semantic — balance structural relevance (this passage is about entities connected to your query) with textual relevance (this passage uses language similar to your question).
The ranked passages, along with entity summaries and graph evidence paths, form the context for answer generation.
Biomedical Research Assistant — Use Case
The Biomedical Research Assistant has been selected as the representative use case for developing and validating ENGRAM because it exercises every capability the system is designed to deliver.
- Dual representation matters because biomedical terminology is full of synonyms: “AD,” “Alzheimer’s Disease,” and “Alzheimer Disease” all refer to the same condition. Hetionet’s entity structure, combined with ENGRAM’s NounPhrase → MAPS_TO resolution and SAME_AS alias edges, handles this naturally. As a result, a query mentioning “AD” activates the same Disease node as one mentioning “Alzheimer’s Disease.”
- Single-step multi-hop retrieval matters because drug-target-pathway-disease chains routinely span three or more hops, across multiple papers. The PPR example above demonstrates exactly this pattern.
- Pattern completion from partial cues matters because researchers often start with incomplete knowledge. A query like “Stanford + synaptic” (i.e., without naming any specific researcher) can activate Thomas Südhof’s work on synaptic transmission and neurodegeneration through the graph connections between institution nodes, biological process nodes, and the passages that cite both.
It is important to note, that the same retrieval pattern applies wherever domain knowledge is scattered across documents that never reference each other directly. Two examples:
- Investment research & due diligence — relevant information lives in SEC filings, earnings transcripts, news articles, and internal research notes. Connections between entities such as companies → suppliers → regulations → geographies can span documents that were never written with each other in mind. A PPR traversal seeded from a company entity can surface supply-chain risks buried in a supplier’s filing three hops away.
- Engineering knowledge bases & incident intelligence — the knowledge exists but is fragmented across incident reports, runbooks, configuration histories, and Slack threads. An incident involving “connection timeout to Redis” should surface similar past incidents → root causes found → configuration changes made → services affected → the engineers who resolved them. Each connection is one graph hop; the full chain spans five or six hops.
I chose biomedical research use case for ENGRAM development because it pushes retrieval requirements the hardest. The Drug → target → pathway → disease chains routinely span 5–7 hops which is exactly where PPR-based retrieval outperforms iterative approaches. Plus, the seed data is freely available: PubMed provides millions of open-access abstracts and articles, while curated knowledge graphs like Hetionet and PrimeKG allow bootstrapping the domain layer and validate resolution quality. To summarise, ENGRAM is designed as a generic GraphRAG method. The biomedical use case simply provided me with the best combination of data availability, multi-hop complexity, and established benchmarks for validation during development.
Query Processing Pipeline
Let’s trace a user query through the system end-to-end. For Part 1, I’ll focus on the retrieval and answer generation path. The multi-agent ENGRAM architecture that enriches responses with pathway analysis, drug interaction checks, and cross-agent synthesis will be covered in Part 2.
Entity Extraction and Resolution
The user types a question in the Research Assistant and optionally selects a source filter: PubMed abstracts only, uploaded PDF documents only, or all sources. The query reaches the Orchestrator, a thin proxy service that manages the query lifecycle without containing domain logic.
The Orchestrator calls the Graph Services Agent which is the central knowledge graph hub to extract named entities from the query (via Claude Haiku) and resolve each one against the knowledge graph (via VoyageAI and ColBERT). Resolved entities are streamed back to the frontend UI immediately as a server-sent event (SSE), so the user sees what the system “understood” from their question while the retrieval pipeline continues working.
Graph Traversal and Passage Retrieval
The resolved entity IDs seed the PPR traversal in Neo4j GDS. The resulting scores identify the most relevant passages across both PubMed abstracts and uploaded documents. This is how a user query about “tau phosphorylation” can surface passages from a paper uploaded by users and relevant PubMed literature in one retrieval step, without the user needing to search each source separately.
Passage ranking blends PPR graph scores with semantic similarity, and the ranked passages are streamed to the frontend UI as another SSE event. The user sees evidence cards appearing in the Passages panel while the answer is still being generated.
Answer Generation
The top-ranked passages, entity summaries, and graph evidence paths are assembled into a context window. Claude Sonnet generates a streaming answer grounded in this evidence. The answer is streamed token-by-token via SSE, so the user sees the response building in real time — there’s no waiting for a complete response before anything appears on screen.
The streaming architecture means the user experience is progressive: resolved entities appear first, then passages, then the answer streams in. Each piece of evidence is visible and can be inspected by user before the final answer is complete.
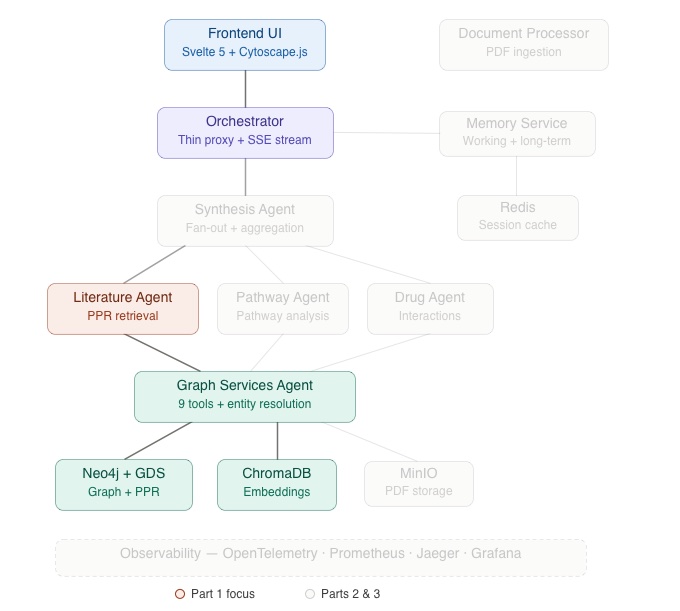
ENGRAM is a multi-agent system. In the architecture diagram above I highlighted data flow relevant for the Part 1. Components of the data flow that Part 1 describes in detail, such as Frontend UI → Orchestrator → Literature Agent → Graph Services Agent → Neo4j + GDS / ChromaDB are shown with full-color coding. Everything covered in Parts 2 and 3 of the blog: Synthesis Agent, Pathway/Drug Agents, Document Processor, Memory Service, Redis, MinIO, and the Observability stack are greyed out.
System Architecture Overview
In contrast, the architecture diagram below shows the full ENGRAM system. Parts 2 and 3 of this series will cover the greyed-out components in detail. I decided to include the full picture here so you can see where the retrieval pipeline fits within the broader system.
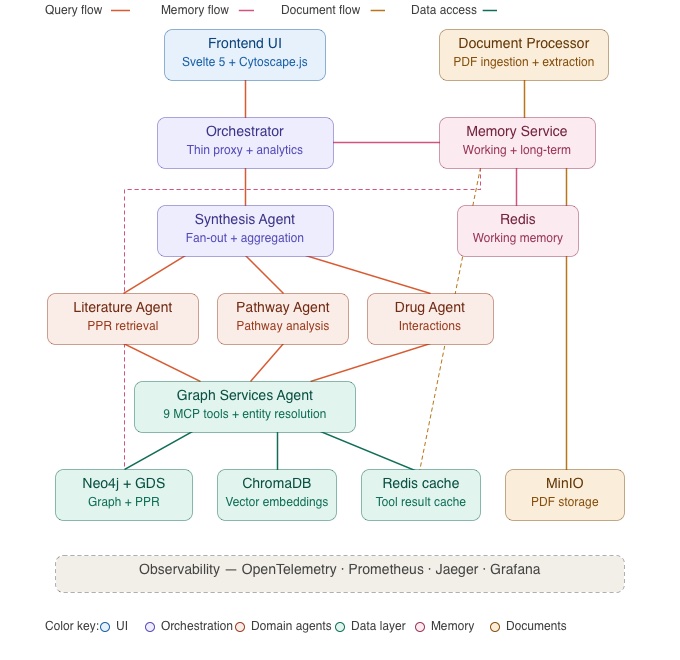
ENGRAM is built as a monorepo of FastAPI micro-services communicating via a lightweight Agent-to-Agent (A2A) protocol which is a simple HTTP task format where one service posts a skill request and receives a structured result. This keeps inter-service communication explicit and easy to debug without the overhead of a message broker. Because each agent runs as an independent service, individual agents can be scaled horizontally based on load e.g., if a deployment handles mostly literature queries, the Literature Agent can run multiple instances behind a load balancer while the Pathway and Drug Agents remain single-instance..
The Graph Services Agent is the central hub. It hosts nine tools (exposed via A2A) for graph operations including PPR traversal, entity resolution, sub-graph extraction, and Cypher queries. All domain agents (Literature Search, Pathway Analysis, and Drug Interaction) communicate with the graph exclusively through this agent rather than connecting to Neo4j directly.
Neo4j with the Graph Data Science library stores the knowledge graph and executes PPR computations. GDS is essential — standard Neo4j doesn’t support Personalized PageRank natively, and PPR is the core retrieval mechanism. ChromaDB handles vector storage for entity embeddings (used in resolution) and passage embeddings (used in semantic ranking). VoyageAI provides the embedding model, and ColBERT (via PyLate) handles token-level reranking for entity resolution precision.
The frontend UI is a Svelte 5 application with Cytoscape.js for interactive graph visualization, Dexie.js for IndexedDB based conversation persistence, and Tailwind CSS 4 with dark / light theme support. It communicates with the backend entirely through SSE streams and REST endpoints (no WebSocket connections).
Each service runs independently via Docker Compose. The monorepo structure keeps shared libraries (A2A clients, embedding utilities, health checks) in a common package while allowing each service to maintain its own dependencies and virtual environment.
Research Assistant UI
The Research Assistant is a three-panel interface designed for exploratory research. Let me walk you through the key functional areas.
Three-Panel Layout
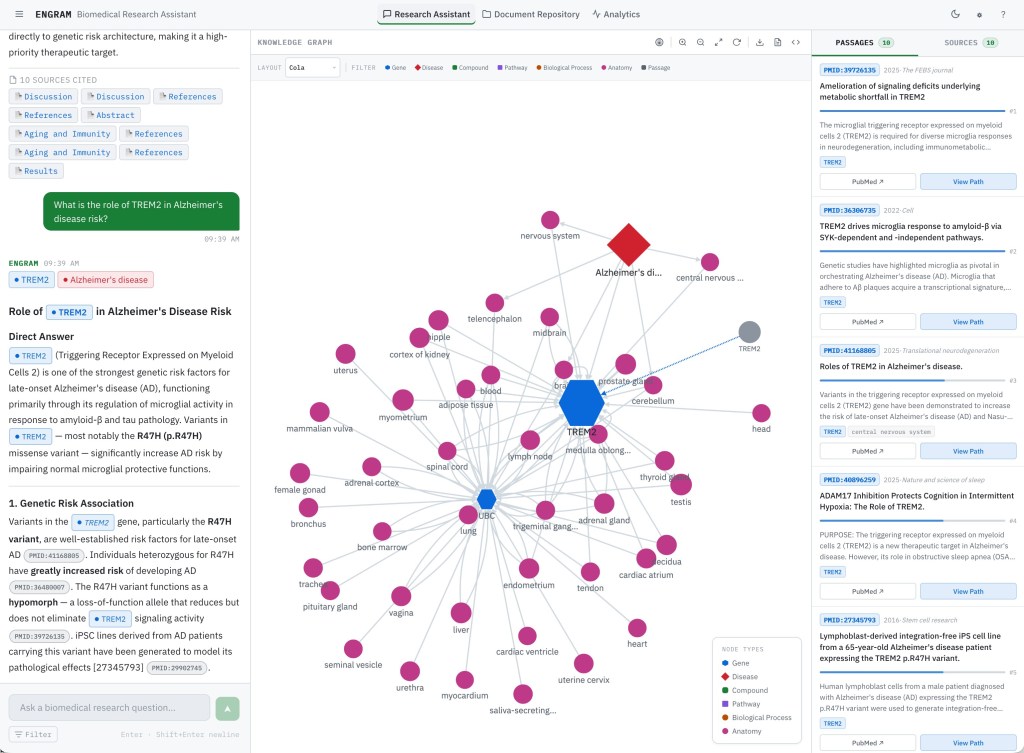
The layout divides the screen into Chat (left), Knowledge Graph (center), and Evidence (right) panels. The Chat panel handles query input and displays streaming responses. The Knowledge Graph panel renders an interactive Cytoscape.js visualization of the PPR-scored subgraph. The Evidence panel shows ranked passages and source documents. All three panels update progressively as a query streams. They’re not sequential steps but parallel views of the same retrieval result.
Asking a Question
The query input sits at the bottom of the Chat panel with a source filter toggle: All Sources, PubMed Only, or Uploaded Documents. If new conversation started, use can see full-screen conversation panel with several sample queries. After submitting a query, the response streams in progressively. First, resolved entities appear showing the user exactly which knowledge graph nodes the system identified from their question. Then the answer text streams in token-by-token. You can see this flow in the video recording below:
Knowledge Graph Visualization
After a query completes, the center panel renders the PPR-scored subgraph. Nodes are color-coded by type, such as diseases, genes, compounds, and pathways. A heatmap overlay indicates PPR activation strength. Brighter nodes received higher relevance scores from the query seeds. Note, you may not see remote nodes dud to low relevance scores assigned by PPR.
The graph is fully interactive as you can see in the video recording below. Clicking a node highlights the passages that reference it in the Evidence panel, creating a direct link between graph structure and textual evidence. Hovering shows entity details. The graph can be zoomed, panned, and exported as SVG or JSON for use in presentations or further analysis.
Passages and Sources
The Evidence panel has two tabs. Passages shows individually ranked passage cards with source badges (PubMed or Uploaded Document), confidence indicators, and expandable full text. Clicking a passage highlights the corresponding entities in the graph. Sources groups passages by their parent document, giving a per-paper view of the evidence which is useful when a researcher wants to see how much evidence comes from a specific study. You can see user flow in the video recording below
Conversation History
Research sessions are persisted to IndexedDB via Dexie.js. The history sidebar lists past conversations, and selecting one restores the full context: graph visualization, passages, and answer text. This means a researcher can run a dozen queries across different topics, close the browser, and return later with every graph and evidence set intact. Note, there is no backend cache dependency.
What’s Next
In this post, we covered ENGRAM’s core retrieval method: how extracted query entities seed a knowledge graph, how Personalized PageRank propagates their relevance across domain relationships then, into literature passages, and finally, how the resulting evidence is ranked and streamed to the user.
But retrieval is only the first stage. In Part 2, I’ll cover how ENGRAM processes uploaded research PDF articles into a unified graph using hierarchical entity extraction (Article → Section → Passage), how a multi-agent architecture fans out queries to specialized domain agents (Literature Search, Pathway Analysis, and Drug Interaction) running in parallel, and how a Synthesis Agent aggregates their findings into a coherent response. I’ll also walk you through the Document Repository UI for managing uploaded papers.
In Part 3, we’ll explore ENGRAM’s hippocampal-inspired memory system which includes working memory in Redis, long-term memory in Neo4j, periodic consolidation that mirrors how the hippocampus transfers memories during sleep, and how memory biases PPR retrieval to improve coherence across multi-turn research sessions.
References
Hetionet — Himmelstein, D. S., Lizee, A., Hessler, C., et al. (2017). Systematic integration of biomedical knowledge prioritizes drugs for repurposing. *eLife*, 6, e26726. Hetionet v1.0 is available under CC0 1.0 Universal (public domain) license at het.io .
PubMed / NCBI — PubMed abstracts accessed via NCBI E-utilities API. National Center for Biotechnology Information, U.S. National Library of Medicine (see pubmed.ncbi.nlm.nih.gov). Usage complies with NCBI’s usage policies and API guidelines .
ENGRAM source code — github.com/pvelua/engram-biomed-research


3 Comments