Part 2 of this blog explains how uploaded research papers join the knowledge graph through hierarchical extraction, and how specialized agents in ENGRAM collaborate to produce richer answers.
Introduction
In Part 1 of this blog series, I described ENGRAM’s core retrieval method: extracting entities from a user query, resolving them to nodes in a biomedical knowledge graph, and using Personalized PageRank (PPR) to spread relevance across graph edges which allowed surfacing connected passages, compounds, and pathways in a single traversal without iterative retrieve → reason → retrieve cycles.
Initially that retrieval pipeline worked with a fixed corpus: ~47K Hetionet entities and ~50K PubMed abstracts loaded at bootstrap. But biomedical researchers don’t just query existing literature. They need uploading their own papers, preprints, and internal reports. These documents need to participate in the same graph-based retrieval as PubMed passages. And the Assistant answers need to go beyond what a single literature retrieval agent can provide e.g., a question about a compound should also surface pathway connections, drug-target relationships, and a cross-agent synthesis that ties everything together.
This post covers the two capabilities that make ENGRAM a complete research system rather than a retrieval demo: long document ingestion into a unified knowledge graph, and a multi-agent architecture where specialized domain agents work in parallel to produce richer, more comprehensive responses.
The Unified Graph Problem
Part 1 showed how PubMed passages connect to Hetionet entities through NounPhrase → MAPS_TO bridges. But PubMed abstracts are short, self-contained texts, typically a single chunk. Research papers are fundamentally different. A 30-page PDF has structure: an abstract summarizing the whole paper, an introduction framing the research question, methods describing experimental procedures, results presenting data, and a discussion interpreting findings. Entities mentioned in the methods section carry different context than the same entities in the discussion.
A naive approach such as chunking the PDF into 300-token passages and treating each one like a PubMed abstract, loses this structural information. A passage from the methods section that mentions “TREM2” in the context of experimental protocol contributes differently to a researcher’s understanding than a discussion section passage that interprets TREM2’s role in neurodegeneration.
ENGRAM preserves document structure through a hierarchical indexing model: Article → Section → Passage. Each level participates in the knowledge graph independently, with entities extracted at every level and linked to Hetionet through the same NounPhrase resolution pipeline used for PubMed. The result is a unified graph where a single PPR traversal retrieves passages from both PubMed abstracts and uploaded documents, ranked by the same blended scoring — without the user needing to search each source separately.
Document Ingestion Pipeline
The architecture diagram below highlights the components involved in document ingestion: the Frontend UI, Orchestrator, Document Processor, and Graph Services Agents plus, the data stores (MinIO, Neo4j, ChromaDB) that participate in the pipeline. The remaining services are greyed out for Part 2 context.
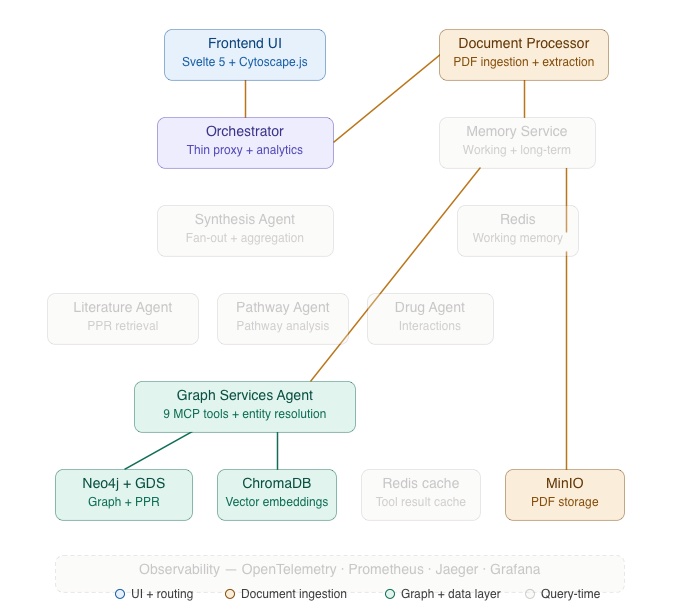
When a researcher uploads a PDF through the Document Repository, it enters a seven-step processing pipeline managed by the Document Processor service.
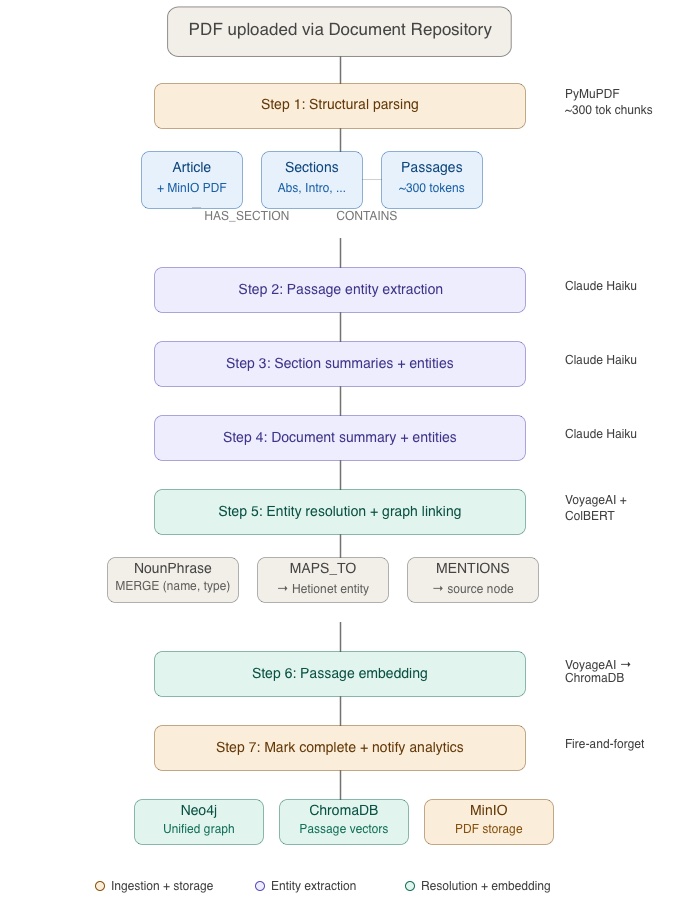
Step 1 – Structural Parsing
The PDF is stored in MinIO (S3-compatible object storage) and an Article node is created in Neo4j with `status: “pending”`. PyMuPDF extracts the text, a section detector identifies structural boundaries (Abstract, Introduction, Methods, Results, Discussion, and other sections), and the text is chunked into passages of approximately 300 tokens. These become a hierarchy of nodes in the graph:
Article (uploaded PDF)
├── Section (Abstract)
│ ├── Passage (chunk 1)
│ └── Passage (chunk 2)
├── Section (Introduction)
│ ├── Passage (chunk 3)
│ ├── Passage (chunk 4)
│ └── Passage (chunk 5)
├── Section (Methods)
│ └── ...
└── Section (Discussion)
└── ...
The graph relationships are explicit: Article -[HAS_SECTION]→ Section -[CONTAINS]→ Passage. This means a query can retrieve individual passages while preserving their structural context as the system knows which section a passage came from and which paper that section belongs to.
Step 2 to 4 – Hierarchical Entity Extraction
Entity extraction happens at three levels, bottom-up:
Passage level. Claude Haiku extracts named entities from each passage individually. A passage about “curcumin inhibiting NF-κB signaling in microglial cells” yields entities like curcumin, NF-κB, and microglia.
Section level. Claude Haiku generates a summary of each section from its constituent passages, then extracts entities from the summary. Section-level extraction captures themes that span multiple passages e.g., an entity might not appear in any single passage but emerges from the section’s overall narrative.
Document level. Section summaries are aggregated into a document summary, and entities are extracted from it. Document-level entities represent the paper’s core topics i.e., the high-level concepts that a researcher would associate with the paper as a whole.
This hierarchical approach ensures that entities are captured at the right level of abstraction. A passage-level mention of “TREM2 expression levels” coexists with a document-level mention of “innate immune response in neurodegeneration”. Both mentions are valid entry points for graph traversal, and PPR will score them differently depending on the query.
Step 5 – Entity Resolution and Graph Linking
Each extracted entity goes through the same resolution pipeline described in Part 1: VoyageAI embedding generates a vector representation next, ChromaDB retrieves the nearest Hetionet candidates using their vector representations generated during initial Hetionet data ingestion, and finally, ColBERT reranking selects the best match. The result is a NounPhrase node with a MAPS_TO edge to a Hetionet entity, and a MENTIONS edge back to the source (Passage, Section, or Article).
The critical design decision here is NounPhrase deduplication. NounPhrase nodes are merged on (name, entity_type)`rather than created fresh per document. This means that when a newly uploaded paper mentions “Alzheimer’s disease,” it doesn’t create a new NounPhrase instead, it reuses the existing one that’s already connected to PubMed passages mentioning the same entity. This cross-document consolidation is what makes the unified graph work: uploaded documents immediately become reachable through the same graph edges that connect PubMed literature.
Step 6 — Passage Embedding
All passages from the uploaded document are embedded using VoyageAI and stored in a dedicated ChromaDB collection (uploaded-passages) with metadata including article ID, section ID, section title, authors, year, and page numbers. At query time, get_passage_rankings() blends PPR scores with semantic similarity across both the PubMed and uploaded collections which makes the retrieval source-agnostic.
Step 7 – Completion
The Article status is set to `”complete”` and a fire-and-forget notification is sent to the Orchestrator’s analytics endpoint to update document processing metrics. The uploaded paper is now fully integrated into the knowledge graph so that its passages, entities, and structural hierarchy can participate in every subsequent query.
Graph Merging and Cross-Document Consolidation
The unified graph only works if entities from different documents are properly reconciled. Two mechanisms handle this.
NounPhrase Deduplication
When the Document Processor extracts “tau phosphorylation” from an uploaded paper, it issues a MERGE on (name: "tau phosphorylation", entity_type: "BiologicalProcess"). If a PubMed passage already created that NounPhrase, the existing node is reused. The new MENTIONS edge simply adds another connection to it. The NounPhrase becomes a shared junction point linking passages from both sources to the same Hetionet entity.
Alias Detection and SAME_AS Edges
Different documents use different surface forms for the same concept: “AD”, “Alzheimer’s Disease”, and “Alzheimer Disease”. The Graph Services Agent includes a merging pipeline that detects these aliases. When multiple NounPhrases map to the same Hetionet entity, bidirectional SAME_AS edges are created using a hub pattern so that the highest-confidence NounPhrase becomes the canonical hub, and all aliases point to it. This avoids O(n²) edges while ensuring that a query mentioning any surface form activates the full cluster.
A ConfidenceScorer aggregates evidence across documents: a MAPS_TO edge that appears in three separate papers receives a higher aggregated_confidence than one from a single paper. This multi-document evidence strengthens the graph over time as more papers are uploaded.
Multi-Agent Architecture
Part 1 described a single-agent query path: the Literature Agent performs PPR retrieval and generates an answer. But biomedical queries often touch multiple knowledge domains. A question like “What are the therapeutic implications of BACE1 inhibition for Alzheimer’s?” involves literature evidence (what papers say about BACE1), pathway analysis (which biological pathways BACE1 participates in), and drug interactions (which compounds target BACE1 and what else they affect).
The full ENGRAM architecture diagram below shows how the domain agents relate to each other and to the shared infrastructure. The query flows from the Orchestrator through the Synthesis Agent, which fans out to the three domain agents. All agents are communicating with the Graph Services Agent for knowledge graph operations.
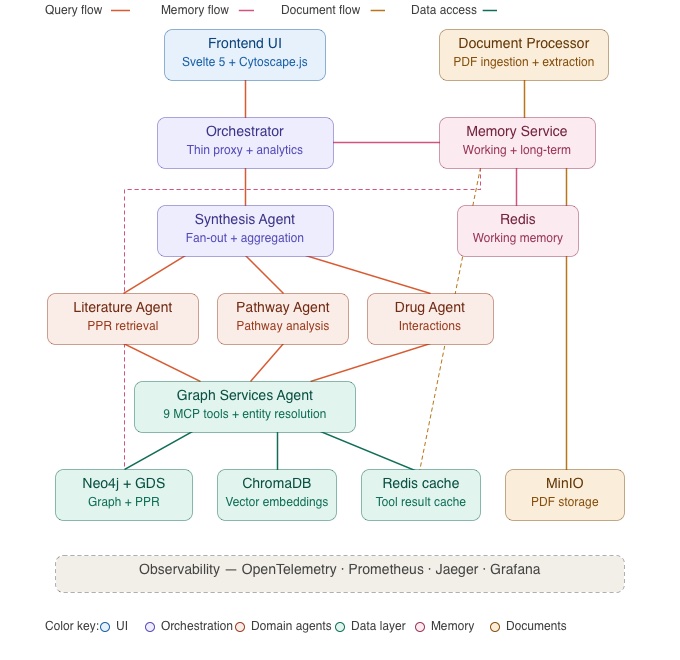
ENGRAM addresses this with a fan-out architecture: the Synthesis Agent examines the resolved entities from the query and dispatches work to specialized domain agents in parallel. The Literature Agent always runs. The Pathway Agent and Drug Agent are activated conditionally based on the entity types present in the query.
Agent Selection
The routing logic is straightforward as it can be driven by the knowledge graph node types that entity resolution produces:
- A resolved Compound entity activates the Drug Agent
- A resolved Pathway or BiologicalProcess entity activates the Pathway Agent
- The Literature Agent always runs, regardless of entity types to find citation sources
A query mentioning both a compound and a pathway activates all three domain agents. A pure disease query activates only Literature. The selection is automatic i.e., the user doesn’t choose which agents to invoke.
The Domain Agents
Each use case specific agent is a FastAPI service communicating via the A2A protocol with the Graph Services Agent. Each has domain-specialized skills that query the knowledge graph from a different angle.
Literature Search Agent performs the PPR-based retrieval described in Part 1: entity extraction, resolution, PageRank traversal, passage ranking, and Claude Sonnet answer generation. It streams the answer token-by-token via SSE, so the user sees the response building in real time. This agent runs for every query and produces the primary answer.
Pathway Analysis Agent traces biological pathways through the Hetionet graph. Its skills include finding shortest paths between entities, listing pathway members (which genes, compounds, or diseases participate in a given pathway), comparing overlap between pathways, and ranking pathways by how many of the query’s entities they contain. When a user asks about TREM2 in the context of neuroinflammation, the Pathway Agent can reveal which signaling pathways connect them.
Drug Interaction Agent analyzes compound-target-disease relationships. It can find direct and indirect targets of a compound (via BINDS and REGULATES edges), identify drug repurposing candidates using PPR-scored disease associations, check for shared targets and pathway conflicts between two compounds, and generate mechanistic narratives explaining how a drug works. When a user asks about curcumin’s therapeutic potential, the Drug Agent surfaces its known targets, related compounds, and potential indications.
Synthesis Agent is the coordinator. It doesn’t query the knowledge graph directly — instead, it fans out work to the domain agents, aggregates their results, and when multiple agents contribute, generates a cross-agent synthesis paragraph using Claude Sonnet that ties the literature evidence, pathway analysis, and drug findings into a coherent narrative.
The Platform Agents & Services
Alongside the domain agents, four platform agents & services provide the shared infrastructure that every query and document operation depends on.
Orchestrator is the system’s front door — every request from the frontend passes through it. It resolves entities from the query, recalls memory context from the Memory Service, proxies the Synthesis Agent’s SSE stream to the frontend, and records analytics metrics to a local SQLite database. Critically, the Orchestrator contains no domain logic. It’s a thin coordination layer: routing, proxying, and bookkeeping. This separation means adding a new domain agent never requires changes to the Orchestrator — only the AgentSelector’s routing table needs updating.
Graph Services Agent is the central knowledge graph hub. It exposes nine tools via the A2A protocol — including PPR traversal, entity resolution, subgraph extraction, and Cypher queries — and is the only service that connects to Neo4j directly. All domain agents access the knowledge graph exclusively through this agent rather than maintaining their own database connections. This single point of access simplifies connection management, enables tool-level caching in Redis, and ensures that graph operations are consistent across agents.
Document Processor Agent handles the PDF ingestion pipeline described earlier in this post: structural parsing, hierarchical entity extraction, entity resolution, passage embedding, and graph linking. It operates independently of the query path — a document can be uploading and processing while the researcher is simultaneously querying the system. After processing completes, a fire-and-forget notification updates the Orchestrator’s analytics metrics.
Memory Service manages ENGRAM’s hippocampal-inspired memory system: working memory in Redis for the current session and long-term memory in Neo4j for persistent knowledge. Every domain agent can store and recall memories through it, and the Orchestrator consults it before every query to inject relevant context. The Memory Service is the focus of Part 3, where we’ll cover its architecture in detail — for now, the key point is that it’s a shared service that all agents communicate with, not a capability embedded in any single agent.
Multi-Agent Query Flow
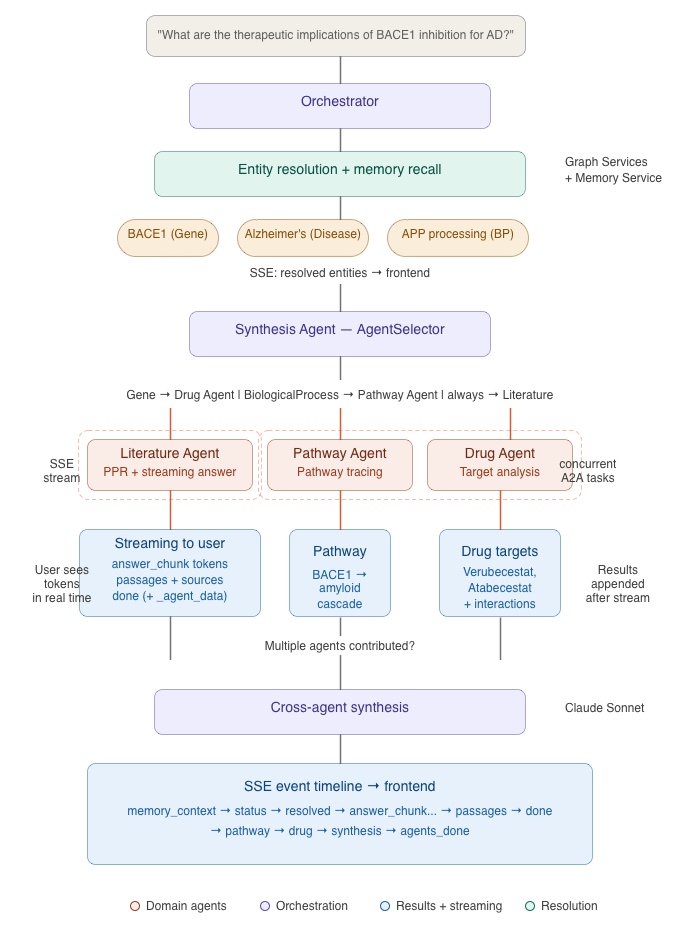
Let me trace what happens when a user submits a query that activates all three domain agents. The example: “What are the therapeutic implications of BACE1 inhibition for Alzheimer’s?
Phase 1 — Entity Resolution and Memory Recall
The Orchestrator receives the query submitted via Frontend UI and calls the Graph Services Agent to extract and resolve entities. “BACE1” resolves to a Gene node, “Alzheimer’s” to a Disease node. Since BACE1 is involved in amyloid precursor protein processing (a BiologicalProcess entity), the resolved entities include types that will trigger all three domain agents.
The Orchestrator also calls the Memory Service to recall any relevant context from the current session (covered in Part 3). Resolved entities and memory context are streamed to the Frontend UI immediately.
Phase 2 — Fan-Out
The Orchestrator forwards the query to the Synthesis Agent, which runs the AgentSelector. The resolved entities include a Gene (BACE1 is associated with Compounds via drug-target edges) and BiologicalProcess types, so the selector activates all three agents: Literature, Pathway, and Drug.
The Synthesis Agent does two things simultaneously:
1. Starts the Literature Agent’s SSE stream so that the user can see answer tokens arriving immediately
2. Launches Pathway and Drug Agents concurrently as background tasks via asyncio.create_task.
This is the key architectural pattern: the user doesn’t wait for all agents to finish before seeing anything. The Literature Agent’s streaming answer appears in real time while the Pathway and Drug Agents work in parallel behind the scenes.
Phase 3 — Domain Agents Results
As the Literature Agent streams its answer (passages, citations, reasoning about BACE1 inhibitors and their clinical history), the Pathway and Drug Agents complete their work independently:
- The Pathway Agent returns the biological pathways connecting BACE1 to amyloid processing and tau phosphorylation, with pathway membership details
- The Drug Agent returns known BACE1 inhibitors (e.g., Verubecestat, Atabecestat), their targets, and drug interaction data
These results are emitted as `pathway` and `drug` SSE events after the Literature Agent’s stream completes.
Phase 4 — Cross-Agent Synthesis
Because multiple agents contributed, the Synthesis Agent generates a synthesis paragraph using Claude Sonnet. This paragraph weaves together the Literature Agent’s evidence about clinical trial outcomes, the Pathway Agent’s mapping of BACE1’s position in the amyloid cascade, and the Drug Agent’s data on specific inhibitor compounds and their target profiles.
The synthesis is streamed as a `synthesis` SSE event, followed by agents_done to mark the end of the multi-agent response.
The SSE Event Timeline
From the user’s perspective, the response builds progressively:
memory_context→ Has context from N memories (if applicable)status→ “Running: Literature, Pathway, Drug agents”resolved→ BACE1 (Gene), Alzheimer’s Disease (Disease)answer_chunk→ Streaming literature answer tokens…passages→ Ranked passage cards appeardone→ Literature agent completepathway→ Pathway analysis resultsdrug→ Drug interaction resultssynthesis→ Cross-agent synthesis paragraphagents_done→ Response complete
Each event updates a different part of the UI: memory context badge in the chat, resolved entities below the query, streaming answer in the chat panel, passages in the evidence panel, and pathway/drug/synthesis results appended to the response.
Architectural Patterns
Several patterns in ENGRAM’s multi-agent design are worth highlighting for readers building similar systems.
A2A Protocol
Inter-agent communication uses a lightweight HTTP task protocol: POST /a2a/tasks {skill, parameters} returns {status, artifacts: [{data: ...}]}. Every domain agent exposes GET /a2a/skills for introspection, making it possible to discover available capabilities at runtime. This is deliberately simpler than a message broker as each agent call is a synchronous HTTP request with a structured response, making debugging straightforward (you can inspect any A2A call with curl).
Fan-Out with Stream Interleaving
The Synthesis Agent’s StreamingAggregator starts domain agents as concurrent asyncio.Task instances before beginning to proxy the Literature Agent’s SSE stream. This means the user sees Literature tokens immediately while Pathway and Drug work happens in parallel. Domain results are appended after the Literature stream completes. The pattern ensures minimal perceived latency — the most important content (the literature answer) streams first, and supplementary analysis arrives as it becomes available.
The _agent_data Sidecar Pattern
The Literature Agent needs to pass internal graph metadata (PPR scores, resolved IDs, passage PMIDs) to the Orchestrator for graph visualization caching, but this data shouldn’t reach the Frontend UI. The solution: the Literature Agent embeds this metadata in its done event as _agent_data. It travels through the Synthesis Agent unchanged. The Orchestrator intercepts it; caches the data for the graph visualization endpoint; and then strips it before forwarding to the Frontend UI. This avoids a separate API call from the Orchestrator back to the Literature Agent.
Thin Orchestrator Principle
The Orchestrator contains no domain logic. It resolves entities, recalls memory context, proxies the Synthesis Agent’s stream, and records analytics i.e., it runs all coordination tasks. Business logic lives in the domain agents. This separation matters as agent count grows: adding a new domain agent requires changes only in the AgentSelector’s routing table and the new agent itself, not in the Orchestrator.
Document Repository UI
The Document Repository is the second top-level tab in ENGRAM’s interface (/documents path in URL). It provides the interface for uploading and managing research papers.
Upload and Processing
Researchers upload PDFs through a drag-and-drop zone or file picker. Each upload immediately appears as a card in the document grid with a processing status indicator. The UI polls the Document Processor for status updates allowing researchers to see the extraction pipeline progressing in real time without leaving the page.
The processing pipeline (described above) runs as a background task. A typical 20-page research paper completes in 2–3 minutes, depending on the number of passages and the entity extraction load. During processing, the document card shows a progress state; once complete, it transitions to a full metadata display.
You can see document upload and processing as well as other use interactions with Document Repository UI in the video recording below.
Document Detail View
Clicking a completed document opens a detail panel showing metadata (title, authors, year, filename), section structure with summaries, extracted entities with their resolution confidence, and extraction statistics (entity count, passage count, processing time). This gives researchers visibility into what the system extracted — useful for validating extraction quality and understanding what graph connections were created.
Search and Filtering
The document list supports text search across titles and filenames, status filtering (processing, complete, error), and sort options. Filters persist across sessions via localStorage. For teams working with large document collections, this makes it practical to find specific papers and check their processing status.
Document Lifecycle Management
Documents can be deleted (removing the PDF from MinIO, nodes from Neo4j, and embeddings from ChromaDB) or reprocessed (re-running the full extraction pipeline). Reprocessing is useful when the entity resolution model improves or when a paper was processed during a period of service degradation.
What’s Next
In this post, we covered how ENGRAM processes uploaded research papers into a unified knowledge graph through hierarchical entity extraction, how specialized domain agents fan out in parallel to produce multi-perspective answers, and how the Synthesis Agent ties their findings together into a coherent response.
But one critical capability is still missing from this picture: memory. In the current description, each query is independent as if the system doesn’t know what the researcher asked five minutes ago. A researcher investigating TREM2’s role in Alzheimer’s across a multi-turn session expects the system to remember their focus and surface increasingly relevant results.
In Part 3, we’ll explore ENGRAM’s hippocampal-inspired memory system which will be the centerpiece of the final post in this series. We’ll cover working memory in Redis that tracks the current session’s entities and discoveries, long-term memory in Neo4j that persists important findings across sessions, a periodic consolidation pipeline that mirrors how the hippocampus transfers memories during sleep, memory-biased PPR retrieval that uses past query context to improve result relevance, and cross-agent memory sharing that lets one agent’s discoveries inform another’s reasoning within the same session.
We’ll close Part 3 with a look at the operational metrics ENGRAM captures such as query performance, entity resolution quality, agent engagement patterns, and memory system health and how the Analytics Dashboard and observability stack (Prometheus, Grafana, Jaeger) make the system’s behavior transparent in production.
References
Hetionet – Himmelstein, D. S., Lizee, A., Hessler, C., et al. (2017). Systematic integration of biomedical knowledge prioritizes drugs for repurposing. eLife, 6, e26726. Hetionet v1.0 is available under CC0 1.0 Universal (public domain) license at het.io.
PubMed / NCBI – PubMed abstracts accessed via NCBI E-utilities API. National Center for Biotechnology Information, U.S. National Library of Medicine. [pubmed.ncbi.nlm.nih.gov](https://pubmed.ncbi.nlm.nih.gov/). Usage complies with NCBI’s usage policies and API guidelines.
ENGRAM source code – github.com/pvelua/engram-biomed-research .
Part 1 – ENGRAM Part 1: GraphRAG with Hippocampal-Like Associative Retrieval.

