How ENGRAM’s agents remember what matters across multi-turn research sessions, and how memory biases retrieval toward coherent results
Introduction
In Part 1 of this series, I introduced ENGRAM’s core retrieval method — spreading query-entity relevance across a knowledge graph via Personalized PageRank. Next, in Part 2 , I showed how uploaded documents join a unified graph and how specialized agents collaborate to produce multi-perspective answers.
But in that description, every query was independent. The system had no way to know that three questions ago, the researcher was investigating TREM2’s role in microglial activation. As a result, when they now ask about “potential anti-inflammatory compounds”, the system can’t prioritize compounds connected to the TREM2 pathway they’ve been exploring. Each query starts from zero.
This is where the hippocampal analogy that inspired ENGRAM’s name reaches its fullest expression. The hippocampus doesn’t just retrieve memories, it consolidates them. Experiences held briefly in short-term circuits are selectively transferred to long-term cortical storage during sleep, with the most important and frequently accessed memories prioritized. Over time, this consolidation shapes which entity associations are strongest, biasing future retrieval toward patterns that have proven relevant.
ENGRAM’s Memory Service implements this same lifecycle: working memory in Redis holds discoveries from the current session, long-term memory in Neo4j persists important findings across sessions, a periodic consolidation pipeline transfers eligible working memories to long-term storage (the ENGRAM’s version of sleep), and memory-biased PPR uses accumulated entity weights to tilt retrieval toward the researcher’s established context.
This post covers the Memory Service architecture, cross-agent memory sharing, memory-biased retrieval, and closes with talking about the operational metrics and Analytics Dashboard that make the ENGRAM system’s behavior transparent.
Why Agents Need Memory
Consider a researcher conducting a multi-turn conversation to investigate into TREM2 and Alzheimer’s disease:
- Query 1: “What is the role of TREM2 in microglial activation?”
- Query 2: “How does TREM2 relate to neuroinflammation pathways?”
- Query 3: “What compounds target neuroinflammation in Alzheimer’s?”
Without memory, Query 3 is a fresh search as. the system has no context that the researcher has been focused on the TREM2/microglia axis. PPR seeds from “neuroinflammation” and “Alzheimer’s” alone, and the results might emphasize general anti-inflammatory compounds rather than those connected to the TREM2 pathway the researcher has been building understanding around.
With memory, Query 3 benefits from everything learned in Queries 1 and 2. The entities explored in previous queries such as TREM2, microglial activation, or specific signaling pathways will carry forward as accumulated weights. When PPR runs for Query 3, these memory-derived weights bias the traversal toward the TREM2-connected neighborhood of the graph, producing results that are coherent with the researcher’s ongoing investigation.
The same principle applies across agents. When the Pathway Agent discovers a TREM2→NF-κB signaling connection in Query 2, that discovery should be available to the Drug Agent when it evaluates compounds in Query 3. Memory enables this cross-agent knowledge transfer.
Memory Architecture
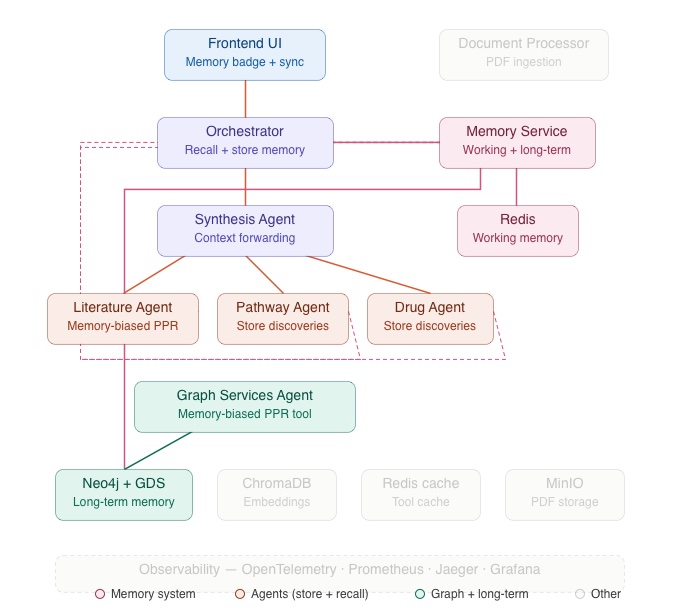
The Memory Service is a dedicated FastAPI micro-service that manages two storage tiers, connected by a consolidation pipeline.
Working Memory — Redis
Working memory captures what’s happening right now. Each query, agent discovery, and Frontend UI interaction generates working memory entries stored in Redis with a session-scoped TTL.
A working memory entry contains a text content summary (what was found or asked), a list of entity IDs linking it to the knowledge graph, the agent that created it (such as Literature, Pathway, Drug, or Frontend), an importance score, and access tracking metadata. Redis keys follow the pattern wm:{session_id}:{memory_id}, with secondary indices for agent-scoped and entity-scoped lookups.
Working memory serves two purposes. First, it provides context for the current session e.g., when the Orchestrator recalls memory before a query, working memories tell the system what the researcher has been exploring in this conversation. Second, it acts as a staging area for consolidation. Working memories that prove important enough are eventually promoted to long-term storage.
Redis’s TTL mechanism ensures working memories naturally expire if the session is abandoned. The default TTL is one hour, configurable per deployment. The eviction policy (allkeys-lru, 512MB cap) prevents memory pressure even under high session volumes.
Long-Term Memory — Neo4j
Long-term memory captures what proved *important* over time. Long-term memories are stored as `(:Memory)` nodes in Neo4j, connected to knowledge graph entities via `[:RELATES_TO {weight}]` edges. This means long-term memories live in the same graph as Hetionet entities, PubMed passages, and uploaded documents which means they can participate in graph traversals.
Each long-term memory node carries an importance score (aggregated from the working memories that produced it), a confidence score (based on how much evidence supported it), a source field (CONSOLIDATION for automatically promoted memories, AGENT for directly stored ones), and provenance tracking (which working memories or agent discoveries contributed to it).
The RELATES_TO edges are the critical mechanism. Each edge carries a weight reflecting how strongly that memory connects to a particular entity. These weights accumulate across sessions e.g., if a researcher repeatedly investigates TREM2, the RELATES_TO weight from their memory nodes to the TREM2 Gene node grows over time. This accumulated weight is what biases PPR in future queries.
Long-term memories are user-scoped and persistent which means they survive across browser sessions, server restarts, and conversation switches. They represent the system’s durable understanding of what this researcher cares about.
Session Identity
Every Frontend UI request includes an X-Session-ID header which is a UUID generated per browser tab and persisted in sessionStorage. This means each browser tab maintains an independent research session with its own working memory context. When a researcher restores a previous conversation from the history sidebar, the Frontend UI re-syncs stored entity IDs to Redis, warming the working memory for the resumed session.
Memory Consolidation — The System’s Sleep Cycle
The hippocampus consolidates memories during sleep by selectively transferring important short-term memories to long-term cortical storage. ENGRAM mirrors this with a periodic consolidation pipeline that runs every 30 minutes (configurable via ENGRAM_MEMORY_CONSOLIDATION_INTERVAL_MINUTES).
The Consolidation Pipeline
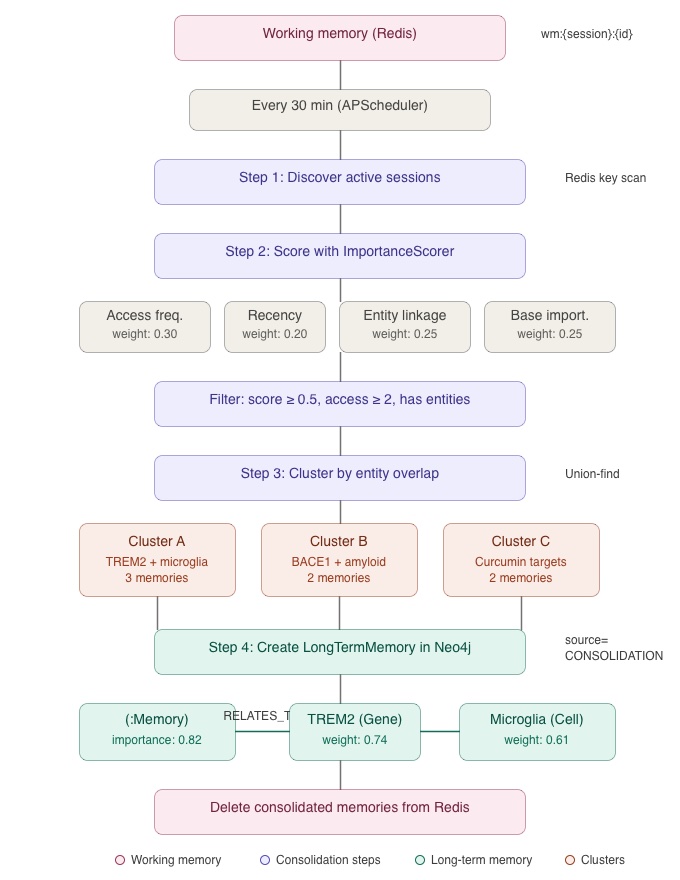
The consolidation job, scheduled via APScheduler, follows a four-step process:
Step 1 — Session discovery. The job scans Redis for active sessions by pattern-matching wm:*:* keys. Each unique session ID represents a research thread with working memories to evaluate.
Step 2 — Importance scoring. For each session, all working memories are recalled and scored by the ImportanceScorer using four weighted factors:
- Access frequency (weight: 0.30): How often was this memory recalled by subsequent queries? Frequently accessed memories are more likely to be relevant.
- Freshness (weight: 0.20): How recently was this memory created or accessed? Recent memories reflect the researcher’s current focus.
- Entity linkage (weight: 0.25): How many knowledge graph entities does this memory connect to? Richly connected memories represent deeper understanding.
- Base importance (weight: 0.25): The inherent importance assigned when the memory was created, based on the significance of the discovery (e.g., a novel drug-pathway connection scores higher than a routine literature citation).
Memories that score below 0.5, have been accessed fewer than twice, or lack entity IDs are filtered out. These thresholds ensure that only memories with demonstrated relevance are promoted.
Step 3 — Entity-overlap clustering. The surviving candidates are grouped using a union-find algorithm: two memories are placed in the same cluster if they share at least one entity ID. This means related discoveries about the same entities are consolidated together rather than stored as separate fragments. A cluster about TREM2/microglial activation might merge three working memories: one from the Literature Agent about TREM2 expression studies, one from the Pathway Agent about TREM2’s signaling pathway, and one from the Frontend UI sync capturing the researcher’s query focus.
Step 4 — Long-term storage. Each cluster becomes a single LongTermMemory node in Neo4j with source=CONSOLIDATION. The node’s importance is the maximum score across cluster members. Its confidence is min(1.0, cluster_size / 5) i.e., larger clusters, backed by more evidence, receive higher confidence. Provenance is preserved in content.source_memories (the list of working memory IDs that contributed). After storage, the consolidated working memories are deleted from Redis.
The result is a progressive distillation: many ephemeral working memories condense into fewer, richer long-term memories that carry aggregated entity weights and cross-agent evidence.
Explicit Session-End Consolidation
Consolidation is also triggered explicitly when a researcher ends a session either by switching to a different conversation in the history sidebar, or closing the browser tab. The Frontend UI checks whether the session meets a consolidation threshold (≥3 working memories or session age ≥10 minutes). If so, it fires a POST /api/memory/consolidation/session/{id}/end request, which triggers consolidation and then clears the session’s working memories from Redis. This ensures no important memories are lost to TTL expiration during natural session transitions.
Memory-Biased PPR — Retrieval Shaped by History
The payoff of the memory system is in how it shapes retrieval. When a query arrives from a researcher with accumulated long-term memories, ENGRAM runs a two-phase PPR instead of the standard single-phase traversal described in Part 1.
How It Works
Phase 1 — Query PPR. Standard Personalized PageRank seeded from the entities extracted from the current query. This is the same traversal described in Part 1.
Phase 2 — Memory PPR. A second PPR traversal seeded from entity IDs that appear in the researcher’s long-term memories, weighted by their `RELATES_TO` edge weights. The Memory Service provides these weights via GET /api/memory/long-term/entity-weights.
Score blending. Both sets of scores are normalized to [0,1] independently, then blended:
final_score = 0.7 × query_score + 0.3 × memory_score
The 0.7/0.3 ratio ensures the current query always dominates i.e., the researcher’s immediate question is the primary signal. But the 0.3 memory contribution nudges results toward the researcher’s established context, surfacing passages and entities that are both graph-relevant to the current query and connected to what the researcher has been investigating across their session history.
The Effect in Practice
Returning to the TREM2 example: when the researcher asks “What compounds target neuroinflammation in Alzheimer’s?” (Query 3), the two-phase PPR produces different results than a memoryless query:
- Without memory: PPR seeds from neuroinflammation and Alzheimer’s. Results include general anti-inflammatory compounds such as curcumin, resveratrol, NSAIDs which are ranked by graph proximity and semantic similarity.
- With memory: The memory phase seeds additional activation from TREM2, microglial activation, and the specific signaling pathways explored in Queries 1 and 2. The blended scores boost compounds that connect to the TREM2/microglia axis e.g., TREM2-modulating antibodies or compounds targeting downstream TREM2 signaling, while still including the general anti-inflammatory results.
The researcher sees results that continue their line of investigation naturally, without having to re-specify their context in every query.
Cross-Agent Memory Sharing
Memory does not need to be private to individual agents. It can be a shared resource that enables knowledge transfer across the multi-agent pipeline described in Part 2.
How Agents Store Memories
Each domain agent stores discoveries as working memories through the Memory Service after completing a task. The Pathway Agent, after tracing a BACE1→amyloid cascade connection, stores a working memory entry with the entity IDs for BACE1, amyloid precursor protein, and the pathway node, plus a text summary of the discovery. The Drug Agent, after finding BACE1 inhibitor compounds, stores its own working memory with compound entity IDs and target relationships.
These stores happen as fire-and-forget operations i.e., they don’t block the response stream. The agent’s primary job (answering the query) is never delayed by memory operations.
How Agents Recall Memories
Before each query, the Orchestrator calls the Memory Service to build an AgentMemoryContext that includes working memories from the current session, long-term memories related to the query’s resolved entities, and aggregated entity weights for PPR biasing. This context is forwarded to every agent that runs.
The Literature Agent uses the entity weights for memory-biased PPR. The Pathway and Drug Agents receive a memory_context_summary (a text summary of relevant prior discoveries) which is prepended to their Claude prompts. This means the Drug Agent answering a question about BACE1 inhibitors might know that the Pathway Agent previously mapped BACE1’s position in the amyloid cascade, enriching its mechanistic reasoning.
The Synthesis Agent takes this a step further: before generating a cross-agent synthesis paragraph, it checks long-term memory for a cached synthesis on the same entity set. If one exists from a previous session, it can build on that foundation rather than starting from scratch.
Shared vs. Private Scope
The Memory Service supports two scopes for memory sharing. Private memories (the default) are visible only to the agent that created them. Shared memories that are created via POST /api/memory/shared/discover, are visible to all agents in the session. Domain agents store their discoveries as shared memories so that cross-agent synthesis benefits from the full picture.
Fault Tolerance — Memory as Enhancement, Not Dependency
A critical design principle: the memory system must never break the query pipeline. All memory operations in the Orchestrator and domain agents are wrapped in try/except blocks. Failures — connection errors, timeouts, unexpected exceptions — are logged at DEBUG level and never propagate to the query handler.
If the Memory Service is down, the Orchestrator emits a memory_context event with has_memory=false and continues with standard PPR (no memory bias). The researcher gets a fully functional response, just without the memory enhancement. The frontend’s memory context badge simply doesn’t appear.
This fault-tolerant design means the Memory Service can be deployed, upgraded, or restarted independently without affecting query availability. Memory is an enhancement layer on top of a fully functional retrieval system.
Frontend UI Memory Integration
The frontend participates in the memory system in two ways.
Memory context display. When the Orchestrator emits a memory_context SSE event with has_memory=true, the chat panel shows a badge: “Context from N memories · PPR biased”. This gives the researcher visibility into when their past session context is influencing results. The badge is hidden on fresh sessions with no memory context.
Session controls. The chat panel provides two session management actions: “Clear Memory” (deletes all working memories for the current session from Redis, resetting to a fresh state) and “End Session” (triggers consolidation, promoting eligible memories to long-term storage, then clearing working memory). These give researchers explicit control over the memory lifecycle.
Background memory sync. After each assistant response, the frontend fires a POST /api/memory/sync/conversation request without waiting for the result. This stores lightweight working memory records from the query’s entity IDs, ensuring that working memory is populated even when domain agents haven’t explicitly stored discoveries. It acts as a safety net i.e., the researcher’s query history is captured regardless of agent behavior.
Operational Metrics and Analytics
Understanding how the system performs requires visibility into every layer: query latency, entity resolution quality, agent engagement patterns, and memory system health. ENGRAM captures these metrics through two complementary systems.
SQLite Analytics Store
The Orchestrator records every query to a local SQLite database (analytics.db) with three tables:
query_eventscaptures per-query timing, entity counts, memory hit counts, and PPR activation metadata.query_agent_spansrecords per-agent timing for each query (auto-populating for new agents without schema changes).document_eventslogs document ingestion completions from the Document Processor.
All writes are fire-and-forget as a result they never block the response stream. The SQLite store is the backend for the Analytics Dashboard described below.
Analytics Dashboard
The Analytics Dashboard is the third top-level tab in ENGRAM’s interface (/analytics in the URL). It provides real-time and historical observability into the full pipeline.
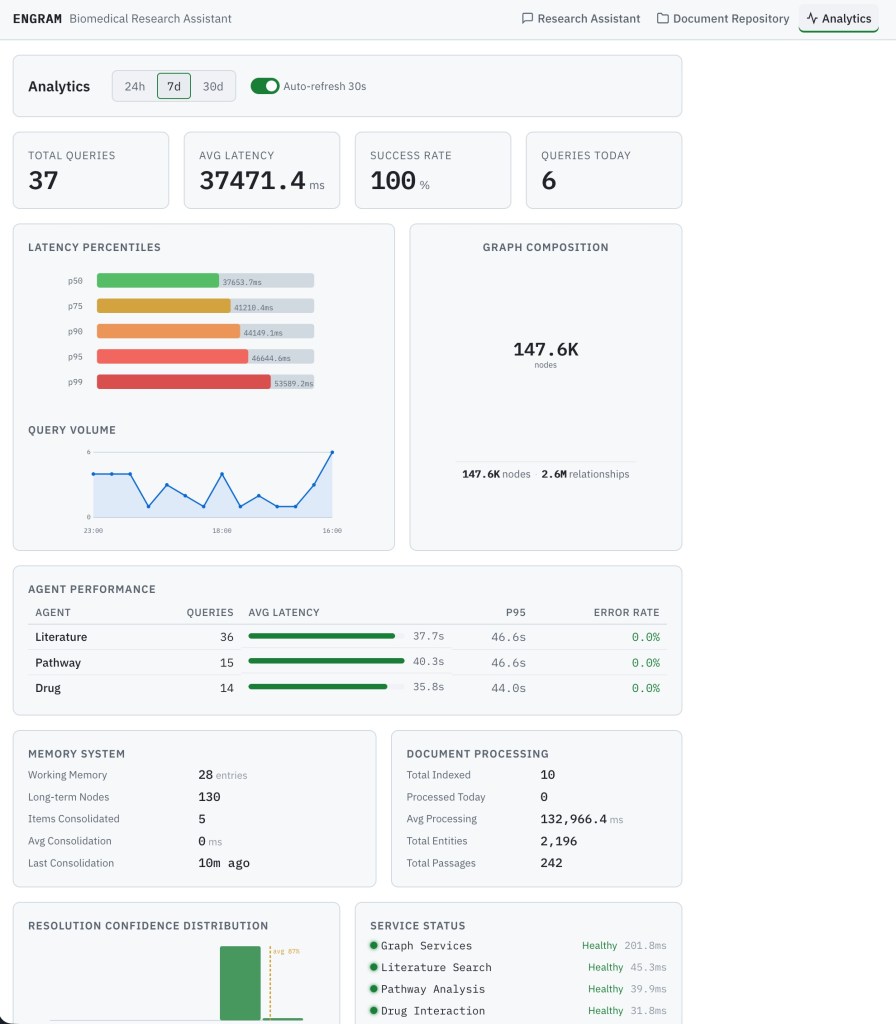
KPI cards show the headline metrics: Total Queries, Average Latency, Success Rate, and Queries Today. These are period-aware as a time-range selector at the top (24h / 7d / 30d) controls all panels, with an auto-refresh toggle that polls every 30 seconds.
Latency percentile chart shows p50 through p99 latency as a bar chart, making it immediately visible whether tail latency is diverging from median performance.
Query volume timeline shows hourly or daily query volume as a line chart with area fill, useful for identifying usage patterns and load peaks.
Agent performance table shows per-agent query count, average latency, p95 latency, and error rate. This panel auto-populates for new agents as a result no configuration needed when a new domain agent is added to the system.
Memory stats panel tracks working memory entries, long-term memory nodes, consolidation runs, and biased PPR usage counts. This is the primary window into memory system health. For example, a drop in consolidation runs or biased PPR usage might indicate a Memory Service issue.
Document processing panel shows total indexed documents, average processing time, and entity/passage counts from the ingestion pipeline.
Confidence histogram displays entity resolution confidence across 10 buckets with an average line, helping identify whether resolution quality is degrading with new document uploads.
Service status panel polls all seven downstream services every 15 seconds and shows a health indicator (healthy / degraded / down) with response latency for each.
Production Observability Stack
Beyond the built-in Analytics Dashboard, ENGRAM emits telemetry to a full observability stack for production monitoring:
OpenTelemetry instruments all services. The HTTPX auto-instrumentation propagates W3C trace context (traceparent header) across every A2A inter-service call, enabling end-to-end distributed tracing.
Prometheus scrapes /metrics endpoints on all eight ENGRAM services every 15 seconds with 30-day retention. Key metrics include engram_ppr_duration_seconds, engram_entity_resolution_duration_seconds, engram_agent_task_duration_seconds, engram_memory_working_entries, and engram_memory_longterm_nodes.
Jaeger provides a distributed trace viewer for inspecting A2A call chains which is useful for diagnosing latency in specific inter-agent calls.
Grafana hosts three pre-built dashboards (System Overview, Query Pipeline, Agent Performance) that visualize the Prometheus metrics. These dashboards are provisioned from configuration files, so they’re reproducible across deployments.
Series Conclusion
Across three posts, we’ve walked you through the full ENGRAM architecture starting from the PPR-based associative retrieval (Part 1), through multi-agent response synthesis and long document support (Part 2), to the hippocampal-inspired memory system and operational observability (this post). The full system is shown in the architecture diagram below.
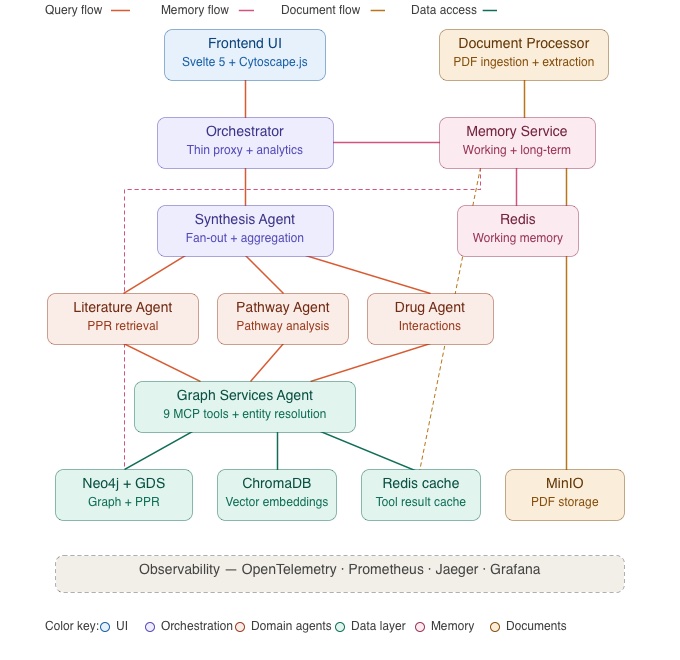
The hippocampal analogy that inspired the project name has threaded through every layer. Pattern completion from partial cues drives the retrieval method. Spreading activation across graph edges replaces iterative LLM reasoning. And the consolidation of working memories into long-term storage (with importance scoring, entity-overlap clustering, and memory-biased retrieval) mirrors the hippocampus’s role in transferring memories from short-term to long-term circuits during sleep.
ENGRAM was designed as a generic GraphRAG method, validated through a biomedical research use case. The same architecture such as a domain knowledge graph seeded with entities relevant to user queries, PPR-based retrieval, hierarchical document indexing, multi-agent specialization, and memory-biased retrieval applies to any domain where knowledge is scattered across documents that never reference each other directly.
One direction we’re actively exploring is whether ENGRAM’s memory architecture can extend beyond domain-specific retrieval to structured memory for general-purpose AI assistants. Current LLM memory systems store user context as flat text summaries with no graph structure. ENGRAM’s approach (entity extraction from conversations, personal knowledge graph construction with deduplication, importance-scored consolidation, and PPR-based context retrieval) could compress conversation history dramatically while selecting more relevant context than recency-based truncation. Early estimates suggest compression ratios of 1000:1 or better from raw conversation history to graph-retrieved context. This is a separate research initiative from ENGRAM Enterprise, but it shares the same core machinery. Stay tuned!
We are also starting work on ENGRAM Enterprise which is a production-grade platform where domain-specific knowledge graphs are constructed from data stored in enterprise databases, internal document repositories, and operational systems. Unlike the biomedical use case where Hetionet provided a pre-built seed graph, Enterprise deployments will use specialized platform agents to analyze enterprise data sources, construct the unified knowledge graph, and generate the domain-specific agents needed for data analysis and recommendations i.e. agents tailored to each organization’s data landscape. The retrieval, multi-agent, and memory capabilities described in this series form the runtime foundation; the platform agents add the ability to build and evolve the domain layer from an organization’s own data. If you’re interested in early access or have a use case that fits the ENGRAM architecture, feel free to reach out.
The full codebase is available at github.com/pvelua/engram-biomed-research on request.
References
Hetionet — Himmelstein, D. S., Lizee, A., Hessler, C., et al. (2017). Systematic integration of biomedical knowledge prioritizes drugs for repurposing. eLife, 6, e26726. Hetionet v1.0 is available under CC0 1.0 Universal (public domain) license at het.io.
PubMed / NCBI — PubMed abstracts accessed via NCBI E-utilities API. National Center for Biotechnology Information, U.S. National Library of Medicine. [pubmed.ncbi.nlm.nih.gov](https://pubmed.ncbi.nlm.nih.gov/). Usage complies with NCBI’s usage policies and API guidelines.
Part 1 — ENGRAM Part 1: GraphRAG with Hippocampal-Like Associative Retrieval
Part 2 — ENGRAM Part 2: Unified Graph Retrieval from Long Documents and Multi-Agent Response Synthesis

